Coat colour basic information (dog)
General information
Mammals inherit two pigments that are the basis for hair colour: eumelanin (black) and pheomelanin (red or yellow). The rich colour varieties seen among dog breeds arise due to genes controlling the amount, extent, and distribution of these two colour pigments. Involved in the production of these pigments in many species including dogs is Melanocortin 1 Receptor (MC1R) gene which is also called Extension. Other genes modify the distribution of these pigments to produce the variety of colours and patterns found in the domestic dog. The Brown gene, Tyrosinase-Related Protein 1 (TYRP1) gene, is a modifier that alters black pigment to brown but does not affect red pigment. Other genes involved in canine coat colour include Agouti (ASIP) which organises the distribution of black and red pigments and Dilute (MLPH) which dilutes black to grey and red to fawn. A lot of other genes specific to some breeds of dogs exist that add white patterns and dilute colours. Below, detailed information on each genetic dog coat colour test offered by LABOKLIN can be found.
Coat colour dog - diagram
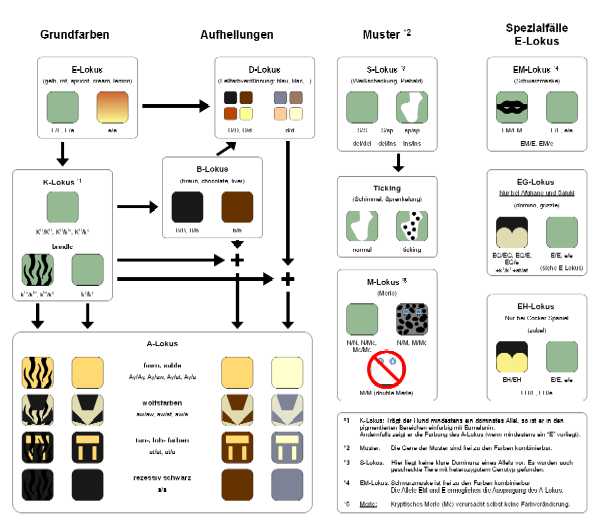
Coat color dog - please click picture for higher resolution
A-locus (agouti)
The A locus is responsible for a number of common coat patterns in the dog. Expression of the A-locus requires any combination of two ky or kbr alleles at the K locus, and at least one E or EM allele at the E locus. The Agouti gene and its variations determine the coat colours fawn / sable (Ay), wild type (aw), tan points ( at ), and recessive black (a). Ay analysis proves absence or presence of the mutation typically responsible for fawn or sable. In fawn / sable dogs this test shows if other agouti alleles are present but hidden (only one copy of Ay). It also demonstrates how many copies of this allele are hidden in dogs, which cannot express agouti types (KB/KB, KB/kbr, KB/ky, at the k locus and/or e/e at the E locus). The analysis of a shows whether a black dog is black due to “recessive black,” or the more common black at the K locus. It also reveals non-black animals carrying “recessive black.” (e.g. German Shepherd Dog, Shetland Sheepdog)
Testinformation A-locus (agouti)
B-locus (coat colour brown, chocolate, livernose,...)
There are two alleles: the dominant full color (B) and the recessive brown (b) which is also known in some breeds as liver, chocolate, sedge, and less frequently, red. Two copies of brown are needed to dilute black pigment to brown. For red or yellow dogs, the brown allele does not dilute the hair color, but will change the colour of nose and foot pads from black to brown if two brown alleles (b/b) are present. This test analyzes whether an animal has 0, 1 or 2 copies of the mutations typically responsible for brown. There are three primary "b" mutations that are responsible for nearly every liver or chocolate dog. A notable exception is the French Bulldog where in addition to these three mutations, there is a fourth cause of chocolate that has yet to be identified.
Testinformation B-locus (coat colour brown,chocolate, livernose,...)
D-locus (coat colour dilution)
The D-locus is the primary locus associated with diluted pigment, which results in coats that would otherwise be black or brown instead showing up as gray, or blue (black), and pale brown or Isabella (brown). The melanophilin gene has recently been shown to be responsible, but not all of the dilute causing mutations have yet been identified. A recessive mutation is the cause of colour dilution phenotypes in the dog. Two alleles (variants) were described: the dominant full colour (D) and the recessive diluted colour (d). Two copies of dilute are needed to lighten black pigment to grey (often called blue), brown pigment to silver and red pigment to cream (also called buff). A diagnostic DNA test identifies the specific variants of the MLPH gene.
Testinformation D-locus (coat colour dilution)
E-locus (coat colour yellow, red, lemon, cream, apricot)
The four alleles (forms) of this gene are black (E), domino/grizzle (EG), melanistic mask (EM) and red (e). The red/yellow coat colour (e) is inherited in an autosomal-recessive trait. E, EG and EM are dominant to e, therefore a dog must have 2 copies of e to be red or yellow. The DNA test offers the detection of carriers. The test does not detect the domino/grizzle (EG) or melanistic mask allele (EM).
Testinformation E-locus (coat colour yellow, red, lemon, cream, apricot)
EG-locus (coat colour domino/grizzle)
The EG allele encodes a coat colour variant called “domino” in Afghan hounds and “grizzle” in Salukis. This pattern is characterized by a pale face with a widow’s peak above the eyes and appears to be unique to these two old dog breeds. EG is dominant over E and e alleles but recessive to the EM allele. Only dogs possesing at/at genotype at the A-locus and ky/ky at the K-locus exhibit this phenotype.
Testinformation EG-locus (coat colour domino/grizzle)
EH-locus* (coat colour sable)
Sable is a coat colour variant in which the expression of eumelanin is partially inhibited. The typical markings are a pale mask and a dark blanket. A mutation at the E-locus, located between E and e, is responsible for this phenotype. It is called eh. A genetic test is only available for American and English Cocker Spaniel. The corresponding coat colour variant is called “grizzle” in Salukis and “domino” in Afghan Hounds. The mutation in these breeds is thought to be in the E-locus as well, however the genetic test is only validated for Cocker Spaniels.
Testinformation EH-locus* (coat colour sable)
EM-locus (melanistic mask allele)
Analysis reveals whether a dog with a mask has one or two copies of this version of the extension locus. Animals with a single copy can produce offspring with or without a mask, while those with two copies will only produce masked offspring. The test may also be applied to black dogs where it may not be possible to tell if there is a mask.
Testinformation EM-locus (melanistic mask allele)
K-locus and K-locus brindle*
The K-locus plays a crucial role in coat colour determination. The findings for it are relative new and changed the understanding of canine colour, as the K-locus includes traits formerly attributed to other genes. KB is the dominant allele, which leads to expression of single coloured fur in pigmented areas. This trait was formerly attributed to AS of the Agouti (A) locus. There are two other alleles, kbr and ky. KB is dominant to both kbr and ky, while kbr is dominant only to ky. kbr is responsible for the brindle pattern. The recessive allele ky allows the basic patterns of the A-locus to be expressed. Same does the kbr allele, but with brindling of any tan, fawn, or tawny areas. Any animal with at least one KB allele will be single-colored in the pigmented areas.
LABOKLIN presently tests for these two alleles. In some breeds, where no brindle is present, this represents a complete analysis of the locus (e.g. the pug). In breeds where the standard disqualifies all but single-colored dogs, testing for these two alleles is sufficient as well. Any animal with two KB alleles cannot produce anything except single-coloured offspring. (e.g. the Labrador Retriever). In breeds where many variations are allowed, these tests can help predict the probability of potential litters to include fawn, sable, tawny, tan point, tricolor or recessive black puppies.
Testinformation K-locus and K-locus brindle*
S-locus (piebald, white spotting)
White spotting in dogs is mostly caused by variations of MITF. Depending on a short interspersed nucleotide element (SINE), the dog is spotted or not. Del is the dominant allele and causes solid or single-coloured dogs (genotype del/del). Dogs that inherit SINE (genotype int/int) homozygous have white markings that either cover at least the ventral surface (mantle pattern) or most of the body (piebald or extreme white spotting). In most breeds, dogs heterozygous for the SINE-insertion (genotype del/int) are solid colored or have minimal white spots, e.g. on the toes. In some other breeds heterozygotes show white undersides, often with a white collar called pseudo-Irish.
Testinformation S-locus (piebald, white spotting)
Interpretation of the test result
Each characteristic is evaluated separately and the genotype for each trait is analysed as follows. Two copies for each characteristic exist in the genome of an animal. Therefore, the outcome are three possible genotypes for either recessive or dominant trait:
Recessive trait:
Genotype N/N: This animal does not inherit the mutation coding for the characteristic. The animal will never pass a mutated allele to it's offspring.
Genotype N/mut: This animal posses one copy of the mutated allele, which makes it a carrier for the trait. It won't exhibit the phenotype. The offspring of the carrier inherit the mutated allele with 50% probability.
Genotype mut/mut: Carrying two mutated copies of the allele, this animal will exhibit the trait phenotypically. Additionally, 100% of it's offspring inherit the mutated allele.
Dominant trait:
Genotype n/n: This animal does not inherit the mutation and does not exhibit the characteristic. The animal will never pass a mutated allele to it's offspring.
Genotype n/Mut: This animal posses one copy of the mutated allele and exhibits the phenotype of the trait. The offspring of the carrier inherit the mutated allele with 50% probability.
Genotype Mut/Mut: Carrying two mutated copies of the allele, this animal will exhibit the phenotype of the trait. Additionally, 100% of it's offspring inherit the mutated allele. Some dominant traits lead to diseases if the animal posses two mutated alleles.
Questiones?
Please contact our molecular biology team for further questions.
LABOKLIN GmbH und Co.KG
Steubenstraße 4
D-97688 Bad Kissingen
Telefon: +49 (0)971 72020
Fax: +49 (0)971 68546
E-Mail: info@labogen.com
Links
back to genetic characteristics
*) Partnerlaboratory